Preclinical Research
Systems Biology
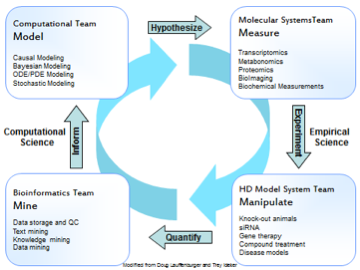 The Systems Biology group’s role is to develop a detailed understanding of Huntington’s disease that can further drug discovery efforts. This includes research to identification of molecular targets or other intervention points, model development, and discovery of predictive biomarkers and outcome measures. Taken together these components allow the basic research effort to nominate therapeutic strategies for the Translational Biology group to prosecute. Our current strategy is based on critical observations from HD patients and is organized into three pillars:
The Systems Biology group’s role is to develop a detailed understanding of Huntington’s disease that can further drug discovery efforts. This includes research to identification of molecular targets or other intervention points, model development, and discovery of predictive biomarkers and outcome measures. Taken together these components allow the basic research effort to nominate therapeutic strategies for the Translational Biology group to prosecute. Our current strategy is based on critical observations from HD patients and is organized into three pillars:
- We know that there is an inverse correlation between (CAG)n and the motoric age of onset. This pillar focuses on the identification of signatures which track with (CAG)n.
- We know that the medium spiny neurons in the striatum seem particularly vulnerable to the mutant huntingtin insult. This pillar focuses on the identification of signatures that vary across cell types.
- We know that HD is a late onset neurological disease that causes a variety of untoward signs and symptoms including disturbances in the circadian cycle. This pillar explores changes in molecular networks and brain circuits that change following mHTT induction and repression to understand causative versus adaptive changes. It also looks at the impact of mHTT on the diurnal cycle.
The Molecular Systems Biology Team within Systems Biology group supports our mission to develop a detailed understanding of the molecular mechanism of HD by focusing on three main areas:
- Molecularly characterize human HD subjects using open, hypothesis free methodologies to generate coherent datasets from multiple biological levels.
- Test hypothesis by perturbing mechanism/pathway using directed methodologies including small molecule, genetic, and viral strategies, in a manner consistent with modeled dysfunction
- Establish and maintain technology platforms to ensure that the rest of the Foundation is empowered with best molecular technologies available. The platforms currently include transcriptomic, proteomic, genomic/epigenomic, and dynamic measures such as metabolomics and lipidomic measures.
Currently ongoing efforts include genome wide association study for HD age-of-onset modifiers, multiple tissue monitoring within each HD and control subject, and transcriptomic & post-translational proteomic characterization of HD mouse brain following with HDAC class2 perturbation.
The HD Model Systems Biology Team within the Systems Biology group supports our mission to develop a detailed understanding of the molecular mechanism of HD as well as supporting preclinical drug testing efforts. Our specific focus is:
- To build cell and animal-based models of HD and deploy for comprehensive phenotyping including biochemical measures, molecular profiling, and behavioral and histochemical measures.
- To identify commonalities and differences between each of the model systems to each other and to the human condition (HD) to allow for more informed choices on how to use the model systems in HD research and development
- To the best of our abilities make the model systems and data widely accessible both within the CHDI collaborative network and to the larger HD research community
The goal of the Bioinformatics Team is to continually grow internal data curation, data management, data mining, data analysis and data delivery capabilities to provide an ever-improving, ever-present Systems Biology context to CHDI project teams and external collaborators. Fundamental to this effort is the adoption – and when necessary, creation – of controlled vocabularies and ontologies that categorize the literature, genomic/genetic data and high-throughput omics assays in a consistent fashion, and permit the semantic integration of public and proprietary data. This, in turn, allows the identification and selection of both comprehensive and precise data subsets by scientists for analyses. Programmatic access to foundational data repositories enable expert computational users the access required to develop high-throughput analytical pipelines over both raw and processed data, and web-based data delivery tools provide non-expert users intuitive access to analytical results.
The Computational Biology Team has the dual responsibility of providing primary analysis of the enormous number of datasets generated within the CHDI network as well as synthesizing those datasets into larger computational models of both Huntington’s disease and of our candidate treatment’s effects on HD. These efforts involve using data from all sources, cellular and stem cell models, animal models across the tree of life, and human patient data. Our goal is to understand the internal wiring of the disease, both molecular and the cellular circuitry of the brain, well enough to make informed predictions on disease progression and potential therapeutics.
Translational Biology
Translational biology refers to the use of biological techniques to enable the discovery of novel therapeutics. Typically, during the drug discovery process, a mechanism or a specific protein becomes a ‘target’ of interest as an intervention point to treat some aspect of the disease. These ‘targets’ need to be specifically impacted by the drugs being developed (either inhibited or activated, for instance), so that we can assess the effect of such drugs in models of the disease. Usually, this process takes place over several years, and it involves a variety of steps to identify molecules with the desired property at the target, either in isolated systems, in cellular systems, and ultimately, in vivo using animal models of the disease. Each one of these steps requires the development of specific assay systems needed to understand that the molecules affect the ‘target’ being sought, and that they do this in ways that are selective and specific (they only modulate this protein target and not others). This type of work entails a very close cooperation between medicinal chemists and biologists, and requires the understanding of the basic biology of the target, and the incorporation of many fields of biology, in order to characterize the drug being developed.
The translational biology group at CHDI is composed of molecular pharmacologists, electrophysiologists, neurochemists and behavioral pharmacologists, and work in the context of a broad network of academic laboratories and contract research organizations (CROs) to design, implement, and evaluate findings of relevance to the development of novel therapeutics. In close collaboration with the chemistry, systems biology and clinical teams, the translational biology team is tasked with developing and implementing methods for assessing the therapeutic relevance of defined drugs or molecular therapies, based on mechanistic hypotheses about the progression of HD, in a variety of domains (behavioral, biochemical, molecular, electrophysiological, or neurochemical) and animal models of relevance to the disease or the mechanism in question.
In some instances, the translational biology team, together with the chemistry team, oversees all aspects of drug discovery from the inception of a program (from the screening phase) to the evaluation of pharmacodynamic and biological effects of the therapeutics in proof of mechanism animal and clinical studies. The translational biology team also supports biological investigations for collaborative work with external pharmaceutical and biotechnology companies, such as Pfizer, GSK, Roche, Astra Zeneca, Lundbeck, Merck, Novartis, Sangamo, Shire, Genzyme, ISIS, Medtronic, and Alnylam. Collaborating CROs include: Brains Online, Psychogenics, Charles River Finland/Cerebricon, Maccine, Evotec, Biofocus, Neuroservice, Ekam, AMRI, and Porsolt.
Chemistry
Medicinal chemists design and synthesize molecules that modulate biological responses. Medicinal chemistry is a key function within the drug discovery process and within the chemistry group at CHDI. All oral medication whether over the counter or prescription were first synthesized by a medicinal chemist. Together with the translational biology team they work together to discover and develop potential therapies. CHDI unlike other not for profit companies has the medicinal chemistry capability to enable the discovery and development of potential therapeutics for Huntington’s disease. Medicinal chemistry also enables the synthesis of research tools that help biologists understand biological pathways and mechanism. This capability also enables the synthesis of high quality compound libraries which are used in biological screens to initiate the drug discovery process.
The chemistry group consist of medicinal chemistry, process research, computational chemistry, chemi-informatics, compound management system, database capture and maintenance, structural biology, drug metabolism and pharmacokinetics (DMPK), bio analytical, formulation, toxicology, and intellectual property. We are responsible for synthesis of validating ligands, design and synthesis of de novo compounds, crystallography of molecular target of interest, evaluation through non-clinical safety in collaboration with the translational biology group.
 CHDI
CHDI